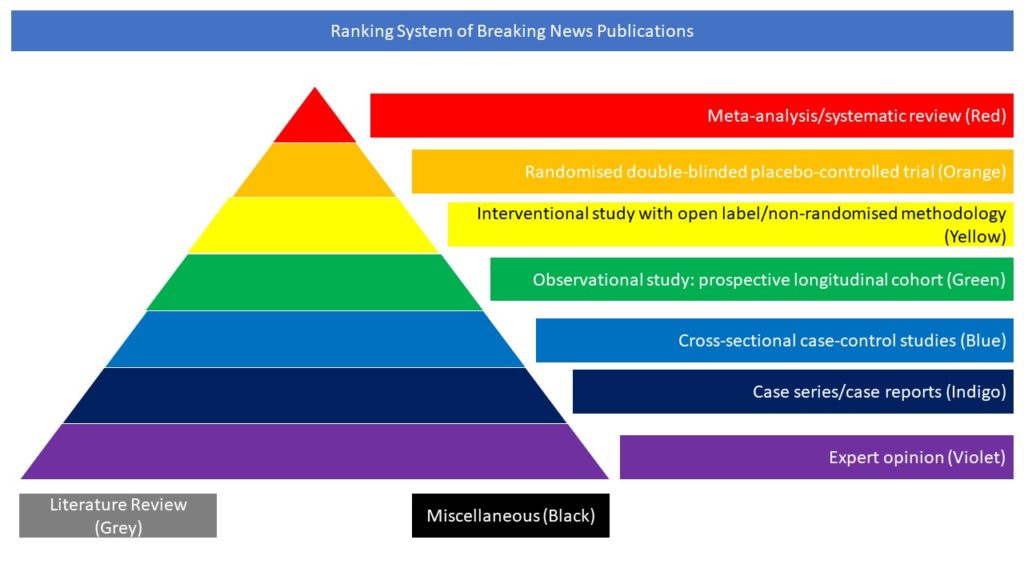
Miscellaneous (Black)
Zoonotic pandemics, such as that caused by SARS-CoV-2, can follow the spillover of animal viruses into highly susceptible human populations. Their descendants have adapted to the human host and evolved to evade immune pressure. Coronaviruses acquire substitutions more slowly than other RNA viruses, due to a proofreading polymerase. The authors of this article found that recurrent deletions in the spike glycoprotein overcame this slow substitution rate. Deletion variants arise in diverse genetic and geographic backgrounds, transmit efficiently, and are present in novel lineages, including those of current global concern. They frequently occupy recurrent deletion regions (RDRs), which map to defined antibody epitopes. Deletions in RDRs confer resistance to neutralising antibodies. By altering stretches of amino acids, deletions appear to accelerate SARS-CoV-2 antigenic evolution and may, more generally, drive adaptive evolution.